
Pseudorabies Resources: Latest PRV Fact Sheet (Updated June 2024) & Additional PRV Resources/Links & Statement
SHIC, launched in 2015 with Pork Checkoff funding, continues to focus efforts on prevention, preparedness, and response to novel and emerging swine disease for the benefit of US swine health. As a conduit of information and research, SHIC encourages sharing of its publications and research. Forward, reprint, and quote SHIC material freely. SHIC is funded by America’s pork producers to fulfill its mission to protect and enhance the health of the US swine herd. For more information, visit http://www.swinehealth.org or contact Dr. Sundberg at [email protected].

Recently, Sun et al. (2021) describe in “Genotype I African swine fever viruses emerged in domestic pigs in China and caused chronic infection” the detection of a second African swine fever virus (ASFV) strain present in two Chinese provinces. The ASF viruses described are genotype 1 viruses, distinct from the currently circulating genotype 2 virus Georgia-07 and its derivatives. These virus isolates (hemadsorption negative) are of lower virulence characterized by a chronic disease presentation including necrotic skin lesions and joint swelling. Data presented suggest the viruses are readily transmissible to contact animals. Notably, pigs infected with these viruses could easily be missed early in a disease outbreak due to their reduced virulence. However, current diagnostic tools PCR (p72-based) or serologic (ELISA-based) should be adequate for detection of infected animals [Sun et al., 2021]. Given their reduced virulence and transmissibility characteristics, it is reasonable to assume these viruses also may be present in other regions of China and Southeast Asia.
The source of these viruses and the nature of their introduction into China is unclear. While they may represent a new introduction of virus from an African source, the striking degree of genetic similarity with NH/P68 and OURT88/3, two genotype I ASFVs isolated in Portugal in the 1960s suggest they may have originated from a European source – possibly imported legally or illegally to be evaluated as potential ASF vaccine candidates in China. Both NH/P68 and OURT88/3 were evaluated as ASF vaccine candidates in the past. Twenty-five to 47% of animals inoculated with a naturally occurring attenuated ASFV isolate, ASFV/NH/P68 (likely a vaccine-derived virus [Portugal et al., 2015]) developed chronic lesions and disease characterized by late fever and viremia and by high levels of anti-ASFV antibodies with marked hypergammaglobulinemia [Leitão et al., 2001]. Less severe post-vaccinal reactions involving fever and joint swelling were described for the ASF live-attenuated vaccine candidate, OUR T88/3 [King et al., 2011].
Currently, ASF vaccines are being developed against Georgia-07 and derivatives (a genotype 2, serogroup 8 virus) [Rock, 2021]. It is highly unlikely these vaccine candidates will provide protection against the viruses (genotype 1, serogroup 4) described here due to the lack of cross protection observed among heterologous ASFV strains.
Given the active Chinese “Belt and Road Initiative”, (https://www.cfr.org/backgrounder/chinas-massive-belt-and-road-initiative) with hundreds of thousands of Chinese working in Africa and many direct flights from African cities to China, there is a possibility that other ASFV strains already may be present or will emerge in China.
Thanks to Dr. Dan Rock, University of Illinois for contributing this article.
King, K.; Chapman, D.; Argilaguet, J.M.; Fishbourne, E.; Hutet, E.; Cariolet, R.; Hutchings, G.; Oura, C.A.; Netherton, C.L.; Moffat, K.; et al. Protection of European domestic pigs from virulent African isolates of African swine fever virus by experimental
immunization. Vaccine 2011, 29, 4593–4600.
Leitão, A.; Cartaxeiro, C.; Coelho, R.; Cruz, B.; Parkhouse, R.M.; Portugal, F.; Vigário, J.D.; Martins, C.L. The non-haemadsorbing African swine fever virus isolate ASFV/NH/P68 provides a model for defining the protective anti-virus immune response. J. Gen. Virol. 2001, 82, 513–523.
Portugal, R.; Coelho, J.; Hoper, D.; Little, N.S.; Smithson, C.; Upton, C.; Martins, C.; Leitao, A.; Keil, G.M. Related strains of African swine fever virus with different virulence: Genome comparison and analysis. J. Gen. Virol. 2015, 96, 408–419.
Rock, D.L. Thoughts on African Swine Fever Vaccines. Viruses 2021, 13(5), 943; https://doi.org/10.3390/v13050943.
Sun et al. Genotype I African swine fever viruses emerged in domestic pigs in China and caused chronic infection. 2021 Emerging Microbes & Infections. https://doi.org/10.1080/22221751.2021.1999779

Evidence suggests African swine fever virus (ASFV) may survive under conditions similar to those observed in transoceanic transport models. In a Swine Health Information Center (SHIC)-funded study, researchers developed a quantitative risk assessment model to estimate the probability that one or more corn or soybean meal ocean vessels contaminated with ASFV would be imported into the US annually. Ultimately, this model can be used to evaluate risk mitigation strategies and critical control points for inactivating ASFV during feed ingredient processing, storage, and transport, and contribute to the design and implementation of biosecurity measures to prevent the introduction of ASFV into the US and other ASFV-free countries. Study authors are Rachel A. Schambow, Fernando Sampedro, Pedro E. Urriola, Jennifer L. G. van de Ligt, Andres Perez, and Gerald C. Shurson.
The published, copyrighted study said the final estimate was conditionally based on five likelihoods: probability of initial ASFV contamination, ASFV inactivation during processing, ASFV inactivation during transport, recontamination, and ASFV inactivation while awaiting customs clearance at US entry. “What if” scenarios were used to explore their impact on risk.
The model estimated complete inactivation of ASFV after soybean extrusion or solvent extraction processes regardless of the initial ASFV contamination probability assumed. The value of recontamination was highly influential on the risk of one ASFV-contaminated soybean meal vessel entering the US. Using the current information available, results showed it would be certain that at least one vessel with ASFV-contaminated soybean meal would be imported once every 21 to ~1,500 years. When all raw corn was assumed to be contaminated and no recontamination was assumed to occur, median probability of one vessel with ASFV-contaminated corn entering the US was 2.02%, or once every 50 years. Days of transport, virus survival during transport, and number of vessels shipped were the most influential parameters for increased likelihood of a vessel with ASFV-contaminated soybean meal or corn entering the US.
The model helped to identify knowledge gaps that are most influential on output values and serves as a framework that could be updated and parameterized as new scientific information becomes available. Researchers propose the quantitative risk assessment model developed in this study be used as a framework for estimating the risk of ASFV entry into the US and other ASFV-free countries through other types of imported feed ingredients that may potentially become contaminated.

Ohio’s Animal Disease Diagnostic Laboratory (ADDL) will now be contributing data to the Swine Disease Reporting System (SDRS) to further enhance capabilities as a surveillance tool and for early detection of pathogens of economic consequence to US livestock production. ADDL provides regulatory testing support for disease control programs and full diagnostic laboratory services for veterinarians, livestock producers and agribusinesses within and beyond Ohio. The SDRS provides data used for disease prevention and biosecurity, disease monitoring, disease management, and disease forecasting. The Swine Health Information Center (SHIC) conceptualized and funded systems for near real-time domestic and global swine disease monitoring to enable better, faster, and more effective response to endemic or foreign infectious diseases. As a result, SHIC helps the industry toward better swine health information to positively impact the long-term sustainability of pork production with the Domestic Swine Disease Monitoring Report.
More than 420,000 tests are performed annually across nine lab sections of the ADDL: aquaculture, avian serology, bacteriology, histology, immunohistochemistry, molecular diagnostics, pathology, serology, and virology. Swine pathogen PCR testing made up approximately 67% of the ADDL molecular diagnostics workload in 2020. Same-day test results are provided by ADDL to clients for PRRS, swine influenza, swine enteric coronaviruses, and Mycoplasma hyopneumoniae. Sequencing is also performed for PRRS and swine influenza to monitor for emerging viruses and epidemiology studies. Ranked 8th nationally in hog and pig inventory, Ohio’s swine industry depends on high quality, fast turnaround testing provided by the ADDL.
The Ohio ADDL is accredited by the American Association of Veterinary Laboratory Diagnosticians (AAVLD), a designation it has maintained since 1999. The lab is also accredited in accordance with ISO/IEC 17025:2017 to perform the following tests: Salmonella enteritidis culture from environmental samples, Salmonella enteritidis testing for eggs, MALDI-TOF for bacterial identification, Pseudorabies gB and gI ELISAs, and whole genome sequencing of bacteria isolates.
At the national level, the ADDL is a Level 1 lab in the USDA National Animal Health Laboratory Network (NAHLN), providing capacity and capabilities for diagnostic testing to respond to emerging and emergency animal disease situations at the state, regional and national level. The laboratory also is one of the FDA Vet-LIRN regional whole genome sequencing centers and is a member of the FDA Global GenomeTrakr network, providing sequencing services for local, national, and international case investigations and surveillance studies.
Key ADDL staff members include Dennis Summers, DVM, DACVPM, Interim State Veterinarian for the state of Ohio, Richard French, MS, DVM, PhD, Laboratory Director, Yan Zhang, MS DVM, PhD, virology section head, and Melanie Prarat, MS. Dr. Summers participated in the inaugural US SHIP House of Delegates meeting earlier this year. Dr. French spent the previous six years building laboratory infrastructure in China and was working in China during the first several years of the ASFV outbreaks. Dr. Zhang identified porcine deltacoronavirus in the US in 2014 and produced the first report linking the virus to a disease outbreak. Prarat previously worked at Plum Island Animal Disease Center studying pathogenesis and vaccine development for classical swine fever virus and coordinates molecular assay development and swine disease epidemiology studies at the ADDL.

There is field and/or experimental evidence that feed and/or ingredients may be potential vectors of African swine fever virus (ASFV) or foot-and-mouth disease virus (FMDV) introduction. And introduction of ASFV or FMDV in a domestic feed manufacturing facility has the potential to unknowingly disseminate those viruses widely. Research is needed to determine optimal methods for decontaminating feed manufacturing facilities, especially equipment that is not designed to be disinfected. The Swine Health Information Center (SHIC) has funded a study, proposed by a group of co-investigators including Dr. Chad Paulk of Kansas State University, to evaluate methods of decontaminating feed manufacturing equipment, using Senecavirus A (SVA), porcine epidemic diarrhea virus (PEDV), and porcine reproductive and respiratory syndrome virus (PRRSV) contamination as domestic, pathogenic surrogates for foreign animal diseases.
Previous research has been conducted to determine the minimum infectious dose of ASFV and FMDV in water and feed (Niederwerder et al. 2019 and Stenfeldt et al. 2021). Recent research has also shown that ASFV and FMDV can survive in various feed ingredients during transboundary, transoceanic shipping conditions (Dee et al., 2021 and Stenfeldt et al. 2021).
Also, field evidence suggests that ASFV can be distributed throughout the feed supply chain (Gebhardt et al., 2021), and this has been confirmed with recent research published from the Feed Safety Team at Kansas State University. Elijah et al. (2021) determined that the distribution of ASFV into the feed manufacturing environment is widespread and persists even after manufacturing additional feed batches initially free of ASFV. This is similar to what is observed with PEDV (Schumacher et al., 2017).
Because ASFV and FMDV can survive in feed during shipping, the US is rightfully concerned that a contaminated feed or ingredient will introduce ASFV or FMDV into the US swine population. Regardless of its method of entry, there is concern that infection of US pigs may result in contamination of the feed supply chain, and rapid and widespread distribution of the virus like what was seen with PEDV, because once ASFV is in a feed mill, it will remain in its environment for an extended period of time.
SHIC continues to look into all routes of entry and dissemination of emerging diseases, not just to identify these pathways, but to do something about them with research of this kind. SHIC is asking other allied feed-related groups to consider contribution to the funding of the project for the benefit of the US swine herd.
Interest in feed biosecurity has been increasing. Recent experimental evidence confirmed African swine fever virus (ASFV), PEDV, Senecavirus A (SVA), and foot-and-mouth disease virus (FMDV) can be transmitted through contaminated feed, providing an avenue for introduction to susceptible pigs via ingestion. One way of reducing the risk of pathogen transmission through feed is to test feed ingredients and feed before they are introduced onto farms and fed to pigs. This would only be possible if sampling and nucleic acid extraction methods would allow efficient detection of pathogens in feed. In a study funded by the Swine Health Information Center (SHIC), principal investigator Dr. Diego Diel, Cornell University, and colleagues focused on comparing the performance of three commercially available nucleic acid extraction kits (CORE, IndiMag, MVP II). Results show the CORE extraction kit outperformed the other two kits evaluated.
These kits were tested on samples spiked with porcine reproductive and respiratory virus (PRRSV), SVA, and PEDV. Feed samples were previously collected as part of a transportation study with PCR testing done in another VDL but offered to this project for further collaboration on adding to PCR extraction comparisons. Overall samples extracted with the CORE kit presented lower Ct value (for PRRSV and SVA) and a higher sensitivity when compared to samples extracted with MVPII or the IndiMag. One of the key issues that remains to be further addressed in future studies is the sampling method to be used for large volumes of feed or feed ingredients.
This initial comparison involved the direct contrast among three extraction methods performed at Cornell University Animal Health Diagnostic Center (MagMax CORE, IndiMag and MVP II). This study was conducted with 28 sample supernatants that had been previously collected and tested for PRRSV, SVA, and PEDV.

The Swine Health Information Center (SHIC)-funded Swine Disease Reporting System (SDRS) initiative has completed another successful year. An aggregated database with diagnostic data from the Iowa State, Kansas State, University of Minnesota, South Dakota State, and Ohio State (beginning in October 2021) veterinary diagnostic labs (VDLs) is regularly updated, with monthly reports and podcasts to SHIC, as well as online interactive dashboards. The database now includes more than unique 950,000 VDL submissions tested by PCR for the five US porcine endemic agents: porcine reproductive and respiratory syndrome virus (PRRSV), porcine epidemic diarrhea (PEDV), porcine deltacoronavirus (PDCoV), transmissible gastroenteritis (TGEV), and Mycoplasma hyopneumoniae. Monthly etiologic summaries of digestive, respiratory, and neurologic diagnostics from the Iowa State VDL are also reported. Interactive online dashboards with filtering capabilities for age category, specimen, geographic region, are kept updated and are available on the project website (https://www.fieldepi.org/sdrs).
The SDRS is the only publicly available source of swine health information from US VDLs. It is also the only source of information on pathogen activity in all age groups (from boar studs to grow-finish pigs). More than 110 monitoring algorithms are currently implemented in the SDRS background to detect changes early in the pattern of agent detection. In addition to SHIC’s website and newsletter, SDRS reports are sent by email to more than 270 registered receivers from 129 organizations/institutions from seven countries (US, Canada, Mexico, Brazil, Chile, Germany, and Spain) and monthly audio and video reports are distributed on the project website, YouTube channel, Swine Cast platform, and LinkedIn.
In July 2020 (SDRS report # 29), monthly PDF reports began to be produced using R Markdown technology in a standard and consistent format. The R Markdown script automates as much as possible the process of generating and updating images, formatting, and description for the monthly detection changes under each agent page. As an ongoing real-time project, there is still the need for staff to interpret the finding, debugging, communicate results with the advisory group, gather feedback, and compile the final report, including the input and feedback from the Advisory group. The usage of an R Markdown script has enhanced SDRS sustainability by reducing the time needed and errors in compiling the final monthly PDF final report.
The SDRS has been built using a building block approach. Initially, SDRS started reporting PRRSV detection from Iowa and Minnesota VDLs and later on added South Dakota, Kansas and Ohio VDLs and additional data for the enteric coronavirus and M. hyopneumoniae. Initially, each agent’s database was kept separate. A restructuring was done to compile, combine, and store all data in a single database. This redesign allowed the database interconnection with the R Markdown to generate monthly reports, analytical tools like R and SAS, and business intelligence tools, like Power BI and Tableau, for data visualization.

The Swine Health Information Center (SHIC) received a call when vesicles were observed in the snout area of pigs on multiple farms in Iowa and Minnesota from January to April 2021. Investigators Jianqiang Zhang, Pablo Piñeyro, and others from Iowa State University (ISU) College of Veterinary Medicine worked on the case. A total of 133 swine vesicular cases with pig ages of three to 6.5 months from Iowa farms were submitted to the ISU Veterinary Diagnostic Lab. All were foot-and-mouth disease virus (FMDV) PCR negative but they were also negative for Senecavirus A (SVA) and other known vesicular viral pathogens, leaving the causative agent(s) unidentified. When standard diagnostic protocols did not reveal satisfying information about the cause, a request for diagnostic fee support was reviewed and approved by SHIC.
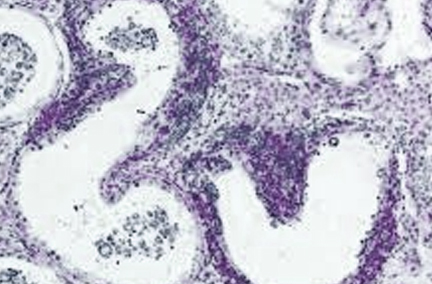
Follow-up investigations on the selected cases were conducted. These cases were also negative for other known vesicular viral pathogens swine vesicular disease virus (SVDV), vesicular stomatitis virus (VSV), and vesicular exanthema of swine virus (VESV). No significant bacteria growth was found from the selected vesicle swabs, suggesting bacteria probably were not the major causative agents. The investigators initiated the SHIC diagnostic assistance request because the known causative viral pathogens of vesicular lesions in pigs – FMDV, SVDV, VSV, VESV, and SVA – each pose a threat to the US swine herd. Currently the US is free of FMDV, SVDV, and VESV; these vesicular disease agents are considered foreign animal disease (FAD) pathogens in the US. VSV is still occasionally reported in the US and SVA is currently endemic in US swine. Complicating diagnostics, the vesicular lesions caused by FMDV, SVDV, VSV, VESV, and SVA are clinically indistinguishable.
Introduction of FMDV would devastate the US pork sector. So early detection and recognition of FMDV is critical to minimize the virus spread and economic burden. However, the endemic infection of SVA may reduce vigilance and result in the assumption that the presence of vesicular lesions is due to SVA so the vesicles would not be reported to State or Federal animal health officials; this would put the early recognition of FMDV at risk.
In summary, no clearly identified infectious causative agent for these vesicular cases has been identified. Nevertheless, the approaches and methods established in the current project can be applied for investigating similar cases in the future studies. It is frustrating for veterinarians, producers, and diagnosticians that the reason(s) for these observed swine vesicular lesions have not been identified although many efforts have been made in regards to sample collections and laboratory testing. However, in order not to miss detecting the true FAD pathogens causing vesicular lesions (e.g. FMDV), when vesicular lesions are observed in pigs, differential diagnosis should still be conducted. In this study, researchers focused on exploring the infectious agents potentially causing vesicular lesion. But, possible non-infectious factors may need to be investigated in the future.

This month’s Domestic Swine Disease Monitoring Report shows a substantial increase in detection of porcine reproductive and respiratory syndrome virus (PRRSV) in the wean-to-market category was associated with a new wave of detection of PRRSV RFLP 1-4-4 L1C variant strain. Also, a moderate increase in PEDV detection in the age category wean-to-market was observed in October. The advisory group has suggested that there may be an opportunity for a national plan to control and eliminate PEDV. Levels of detection of M. hyopneumoniae by PCR are at expected levels for this time of year. In the podcast, the SDRS hosts talk with Dr. Brigitte Mason, health assurance veterinarian from Country View Family Farms, about her experience with animal health management, disease management, control, and her advice to the swine industry to better handle animal health interventions.

In the November Global Swine Disease Monitoring Report, read about African swine fever (ASF) in the Dominican Republic where authorities keep registering new disease reports. So far, over 283 reports have been confirmed in the island nation. ASF is spreading westward in Germany; ASF appeared 43.5 miles (70 km) west of previous reports. In Russia, new ASF outbreaks in large pig farms, including a new site of the country’s largest pork producer, are detailed. Classical swine fever (CSF) has been diagnosed in Brazil outside the free zone, the first confirmation since 2019.